Yugant Yugant
A Unified Framework for 2D-3D and 3D-3D Image Registration2D-3Dおよび3D-3D画像レジストレーションのための統一フレームワーク
This project presents a unified computational framework designed to handle two fundamentally different but related image registration problems: the alignment of a 3D model to a 2D image (2D-3D registration) and the alignment of two 3D volumetric datasets (3D-3D registration). Image registration is a foundational operation in medical image analysis, underpinning applications ranging from intraoperative surgical navigation and implant positioning to the analysis of molecular structures in electron microscopy. The development of a single, cohesive framework that can address both registration types is motivated by the observation that many of the core computational components — such as the optimization strategy, the rendering pipeline, and the similarity metric — can be shared or adapted across both tasks.
本研究は、三次元モデルを二次元画像に位置合わせする2D-3Dレジストレーションと、二つの三次元ボリュームデータセットを位置合わせする3D-3Dレジストレーションという、根本的に異なるが関連する二つの画像レジストレーション問題を統合的に扱う計算フレームワークを提示します。画像レジストレーションは医用画像解析の基盤的操作であり、術中外科ナビゲーション・インプラント位置決めから電子顕微鏡での分子構造解析まで幅広い応用を支えています。最適化戦略・レンダリングパイプライン・類似度指標などの中核的な計算コンポーネントが両タスク間で共有・適応できるという観察が、統一フレームワーク開発の動機となっています。
Focus研究焦点
A Unified Framework for 2D-3D and 3D-3D Image Registration2D-3Dおよび3D-3D画像レジストレーションのための統一フレームワーク
This project presents a unified computational framework designed to handle two fundamentally different but related image registration problems: the alignment of a 3D model to a 2D image (2D-3D registration) and the alignment of two 3D volumetric datasets (3D-3D registration). Image registration is a foundational operation in medical image analysis, underpinning applications ranging from intraoperative surgical navigation and implant positioning to the analysis of molecular structures in electron microscopy. The development of a single, cohesive framework that can address both registration types is motivated by the observation that many of the core computational components — such as the optimization strategy, the rendering pipeline, and the similarity metric — can be shared or adapted across both tasks.
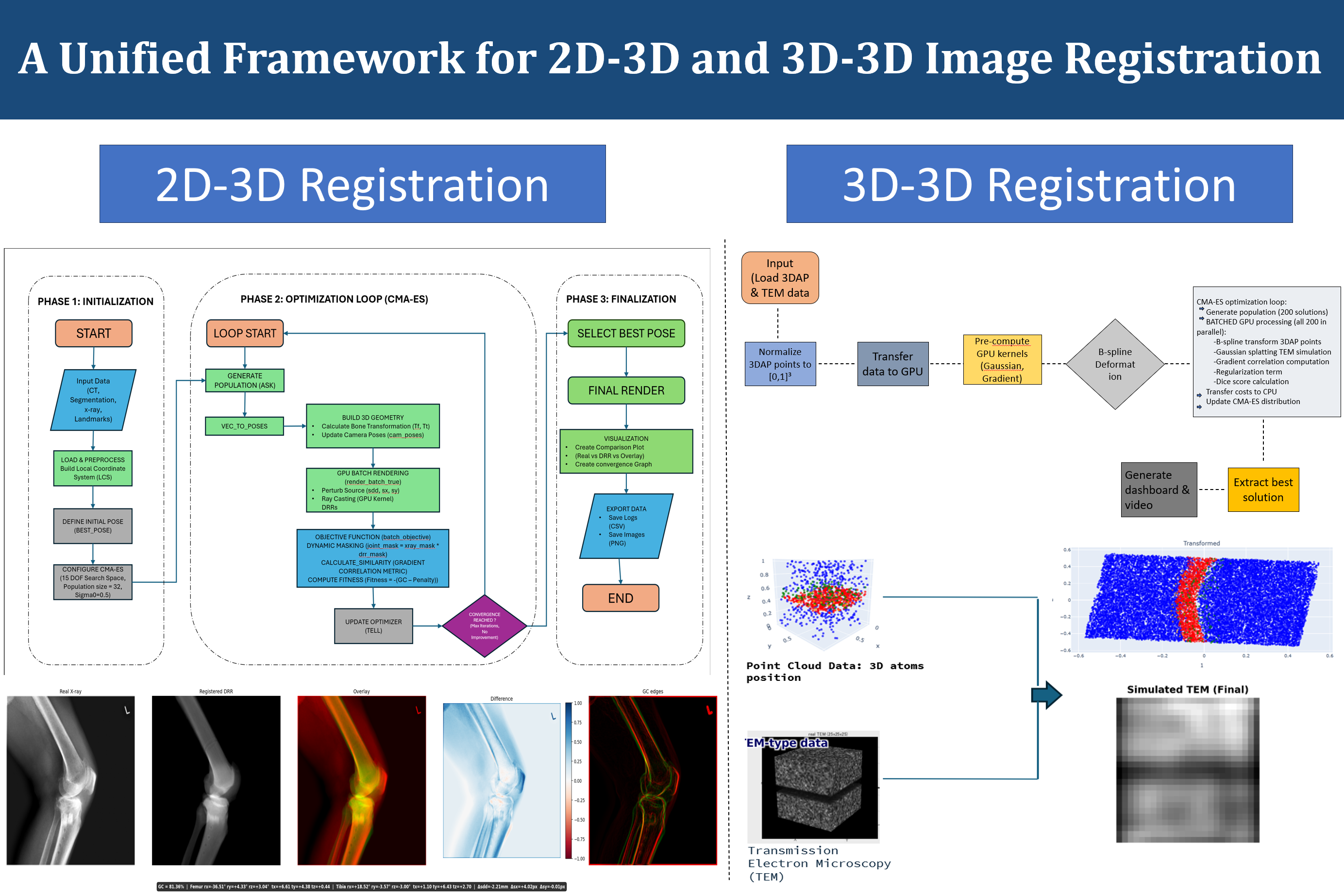
The 2D-3D registration component addresses the classical problem of aligning a three-dimensional bone model to a real X-ray image. This is a critical step in many orthopedic applications, including fluoroscopy-based surgical guidance and the computation of implant kinematics from postoperative radiographs. The framework structures the 2D-3D registration process into three distinct phases. In the initialization phase, the input data — consisting of CT segmentation, X-ray image, and optionally anatomical landmarks — is loaded and preprocessed to establish a local coordinate system and define an initial pose estimate. The optimization phase employs CMA-ES (Covariance Matrix Adaptation Evolution Strategy), a derivative-free black-box optimization algorithm well-suited for non-convex, high-dimensional problems.
Within the CMA-ES loop, candidate bone poses are evaluated by rendering batched digitally reconstructed radiographs (DRRs) using GPU-accelerated ray casting. For each candidate pose, a 3D bone geometry is assembled from the CT segmentation, projected using the known X-ray imaging geometry, and compared against the real X-ray image using a gradient correlation metric combined with a dynamic masking strategy. The dynamic masking restricts the similarity computation to a joint mask derived from the X-ray image and the current DRR, focusing the optimization on anatomically relevant regions. A penalty term is also included in the fitness function to discourage implausible poses.
The 3D-3D registration component targets a different application domain: the alignment of molecular point cloud data obtained from 3D Atom Probe (3DAP) tomography with simulated Transmission Electron Microscopy (TEM) images. In this setting, the registration problem involves finding the spatial transformation that best aligns the point cloud representation of atomic positions with the volumetric TEM simulation data. The optimization strategy again employs CMA-ES, but the similarity metric is adapted for 3D-3D comparison, using B-spline deformable transformations, Gaussian splitting for TEM simulation, and gradient correlation for assessing alignment quality. GPU-based batch processing of all candidate solutions enables efficient exploration of the search space.
The unification of both registration frameworks under a common implementation provides significant practical advantages. Shared components — such as the CMA-ES optimizer, the GPU batch processing pipeline, and the output generation modules (logs, CSV exports, comparison plots, and convergence graphs) — reduce code duplication and facilitate future extensions to new imaging modalities or registration problems. The finalization phase in both cases selects the best pose from the optimization run, generates visualization outputs, and exports the registration results in standardized formats. This framework represents a versatile and extensible approach to image registration that bridges applications from intraoperative orthopedic surgery to nanoscale materials science.
本研究は、三次元モデルを二次元画像に位置合わせする2D-3Dレジストレーションと、二つの三次元ボリュームデータセットを位置合わせする3D-3Dレジストレーションという、根本的に異なるが関連する二つの画像レジストレーション問題を統合的に扱う計算フレームワークを提示します。画像レジストレーションは医用画像解析の基盤的操作であり、術中外科ナビゲーション・インプラント位置決めから電子顕微鏡での分子構造解析まで幅広い応用を支えています。最適化戦略・レンダリングパイプライン・類似度指標などの中核的な計算コンポーネントが両タスク間で共有・適応できるという観察が、統一フレームワーク開発の動機となっています。
2D-3Dレジストレーションコンポーネントは、三次元骨モデルを実X線画像に位置合わせする古典的な問題に取り組みます。これは透視X線ベースの外科ナビゲーションや術後X線からのインプラント動態計算など、多くの整形外科応用において重要なステップです。フレームワークは2D-3Dレジストレーションプロセスを三つの段階に構造化します。初期化段階では入力データ(CTセグメンテーション・X線画像・オプションの解剖学的ランドマーク)を読み込み前処理して局所座標系を確立します。最適化段階では非凸・高次元問題に適した微分不要のブラックボックス最適化アルゴリズムCMA-ESを採用します。
CMA-ESループ内では、GPU加速レイキャスティングを使用したバッチDDR(デジタル再構成X線画像)レンダリングにより候補骨姿勢を評価します。各候補姿勢でCTセグメンテーションから三次元骨形状を組み立て、既知のX線撮像幾何学で投影し、動的マスキング戦略を組み合わせた勾配相関指標で実X線画像と比較します。動的マスキングはX線画像と現在のDRRから導出された結合マスクに類似度計算を制限し、解剖学的に関連する領域に最適化を集中させます。
3D-3Dレジストレーションコンポーネントは異なる応用領域を対象とします。三次元アトムプローブ(3DAP)断層撮影から得られた分子点群データとシミュレーション透過電子顕微鏡(TEM)画像の位置合わせです。最適化には再びCMA-ESを採用しますが、類似度指標はBスプライン変形変換・TEMシミュレーションのためのガウス分割・アライメント品質評価のための勾配相関を使用した3D-3D比較に適応しています。全候補解のGPUバッチ処理により探索空間の効率的な探索が可能です。
共通実装下での両レジストレーションフレームワークの統一は重要な実用的利点を提供します。CMA-ES最適化器・GPUバッチ処理パイプライン・出力生成モジュール(ログ・CSVエクスポート・比較プロット・収束グラフ)などの共有コンポーネントがコード重複を削減し、新しい撮像モダリティやレジストレーション問題への将来の拡張を容易にします。このフレームワークは術中整形外科手術からナノスケール材料科学まで応用が架け橋となる多目的で拡張可能なアプローチを表しています。
Research Classification Overview
The following table categorizes each research project according to its primary methodological and clinical research classes.
Project
Class 1
Class 2
Class 3
Motion Analysis of Ballet Movements (Arisa)
Motion Analysis
Multimodal Learning
Posture Evaluation
Pelvic Tilt Estimation (Daisuke)
Depth Map Estimation
Landmark Detection
2D→3D Reconstruction
Survival Prediction in Fulminant Myocarditis (Itoi)
Foundation Model Transfer
Multimodal Learning
Survival Prediction
Swallowing Movement via 4DCT (Izumi)
Segmentation
Spatiotemporal Analysis
Clinical Quantification
ONFH Automated Classification (Mamoru)
Segmentation
Automated Classification
Multi-scheme Diagnosis
AP Depth Estimation from X-ray (Naoaki)
Depth Map Estimation
2D→3D Reconstruction
Vertebral Analysis
BMD Age Analysis with CT (Natsu)
CT Analysis
Bone Density Analysis
Population Study
Internal Structure from Surface Scan (Ryota)
Statistical Shape Model
Surface-to-Bone Estimation
3D Reconstruction
Sex/Age BMD Analysis with CT (Uwai)
CT Analysis
Sex-stratified Analysis
Bone Density Analysis
Hand Bone 2D–3D Reconstruction (Yanis)
Depth Map Estimation
2D→3D Reconstruction
Surgical Planning
Unified Registration Framework (Yugant)
Image Registration
2D-3D Registration
3D-3D Registration
Class Taxonomy Definitions
2D→3D Reconstruction: Methods that recover three-dimensional anatomy from one or more 2D images using depth estimation, back-projection, or learned volumetric inference.
Automated Classification: Systems that assign patient data to predefined clinical categories (e.g. disease severity stages) without human intervention.
Bone Density Analysis: Quantitative extraction and analysis of cortical and/or trabecular mineral density from CT or DXA imaging.
CMA-ES Optimization: Use of the Covariance Matrix Adaptation Evolution Strategy for black-box, gradient-free optimization in registration or pose estimation.
Clinical Quantification: Extraction of objective, numerical measurements from medical images to characterize physiological or pathological processes.
CT Analysis: Processing and analysis of computed tomography volumes for segmentation, density measurement, or 3D modeling purposes.
Depth Map Estimation: Prediction of pixel-wise depth (distance from detector or reference plane) from 2D medical images, typically X-rays.
Foundation Model Transfer: Application of large models pre-trained on broad data corpora to new, often narrower clinical domains.
GPU Rendering / DRR: Real-time generation of digitally reconstructed radiographs via GPU ray casting, used in 2D-3D registration.
Image Registration: Spatial alignment of two or more images or models, encompassing both 2D-3D and 3D-3D settings.
Landmark Detection: Localization of anatomically defined keypoints in images for geometric reference or downstream computation.
Motion Analysis: Quantification of body segment kinematics from motion capture or dynamic imaging data.
MRI Segmentation: Delineation of anatomical or pathological structures in MRI volumes using deep learning networks.
Multimodal Learning: Integration of heterogeneous data streams (images, text, motion, clinical variables) within a single learning framework.
Musculoskeletal Modeling: Construction of subject-specific biomechanical models incorporating bone, muscle, and joint geometry.
Non-invasive Imaging: Approaches that infer internal anatomy from surface measurements without radiation or invasive procedures.
Population Study: Statistical characterization of imaging biomarkers across a defined cohort stratified by age, sex, or clinical group.
Posture Evaluation: Automated scoring or assessment of body posture quality based on motion, anatomy, and expert criteria.
Segmentation: Delineation of specific structures or regions within medical images using deep learning or classical methods.
Spatiotemporal Analysis: Joint analysis of spatial and temporal dimensions in dynamic imaging data to characterize motion or physiology.
Statistical Shape Model (SSM): Compact representation of shape variability across a population, used for shape prior regularization and fitting.
Survival Prediction: Estimation of patient survival probability from clinical and/or imaging data using machine learning models.
Surgical Planning: Use of computational models or reconstructions to support preoperative decision-making and implant selection.